Mass Spectrometry (LC-MS/MS) & Proteomics Services
ITSIBIO »
High‑Resolution LC‑MS/MS Analysis for Confident Protein Characterization
Our mass spectrometry services deliver fast, high‑quality LC‑MS/MS analysis for proteins and complex biological samples. We combine advanced instrumentation with expert scientific support to generate sensitive, specific, and publication‑ready data with rapid turnaround times.
Reach out to us for more information. Also, click a button below to view our list of citations involving our LC-MS-MS services or to fill out a project submission form to receive a quote. You can read more about the services we offer below.
Protein Identification – (S50)
Read More
Protein Relative Quantitation by TMT Multiplexing – (S90)
Read More
Label Free Quantitative Proteomics – (LFQ)
Read More
Post Translational Modification (PTM) Analysis – (S60)
Read More
Our LC MS/MS Workflow includes
- Sample Preparation: Reduction, alkylation, and tryptic digestion using automated or manual workflows
- LC‑MS/MS Analysis: Nano‑LC separation and high‑resolution mass spectrometry
- Data Analysis: Protein identification using established proteomics software and curated databases
- Reporting: Comprehensive reports including identified proteins, peptides, quantitative results, and raw data access
Typical turnaround time: 2–3 weeks for most services
Why Choose Our Mass Spectrometry Services?
- High‑resolution LC‑MS/MS platforms
- Comprehensive proteomics workflows
- Advanced bioinformatics analysis
- Experienced scientific team
- One‑stop sample‑to‑report service
1. Protein Identification – (S50)
Accurate Protein Identification by LC‑MS/MS
We provide high‑sensitivity protein identification services for academic, biotech, and pharmaceutical research. Using advanced LC‑MS/MS workflows, we support the analysis of purified proteins, gel bands, protein mixtures, and complex biological samples with exceptional accuracy and reproducibility.
Gel‑Based Protein Identification
- Identification from SDS‑PAGE gel bands or gel spots
- Compatible with most gel matrices and stains
- If you use silver stain use an MS compatible stain.
Protein Identification in Solution
- Suitable for purified proteins or complex mixtures of proteins.
- We can work with dilute as well as concentrated samples.
- ITSI will perform a buffer exchange if the protein solution is incompatible.
2. Protein Relative Quantitation by TMT Multiplexing – (S90)
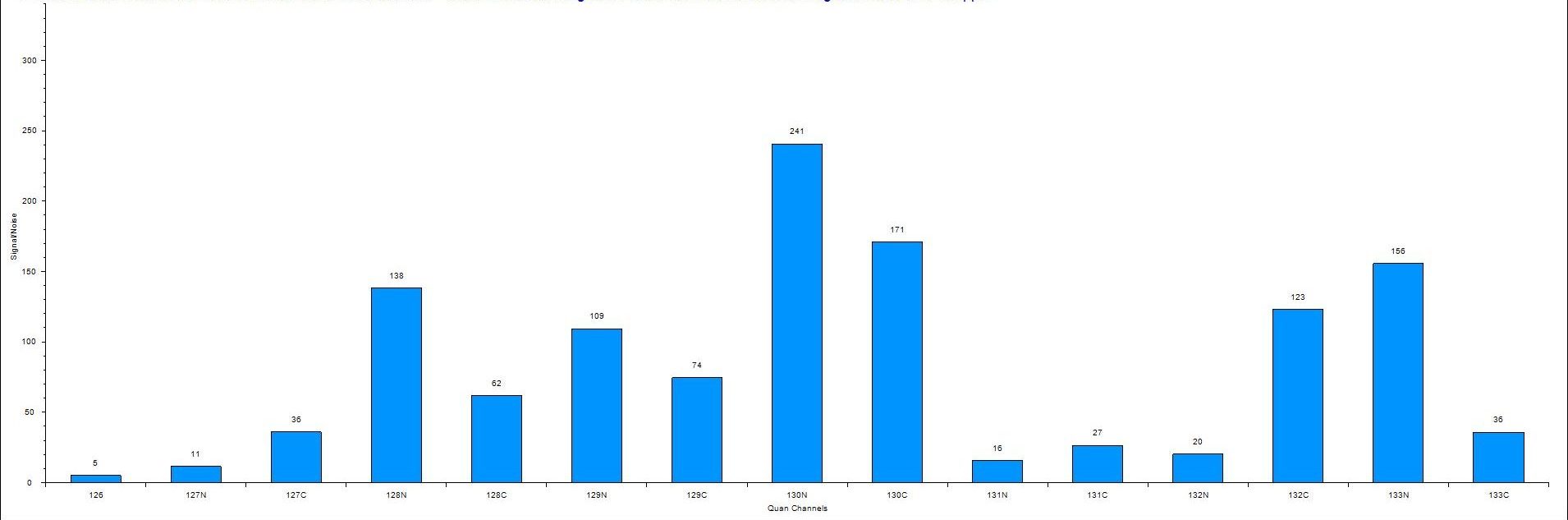
High‑Precision, Multiplexed Quantitative Proteomics
We offer label‑based quantitative proteomics using multiplexed Tandem Mass Tag (TMT) strategies to measure relative protein abundance across multiple experimental conditions in a single LC‑MS/MS run. This approach provides high precision, excellent reproducibility, and robust comparative analysis.
Key Advantages
- Simultaneous comparison of multiple samples
- Reduced technical variability
- Ideal for comparative proteomics, biomarker discovery, and drug research
- High‑resolution LC‑MS/MS ensures confident protein identification and quantitation
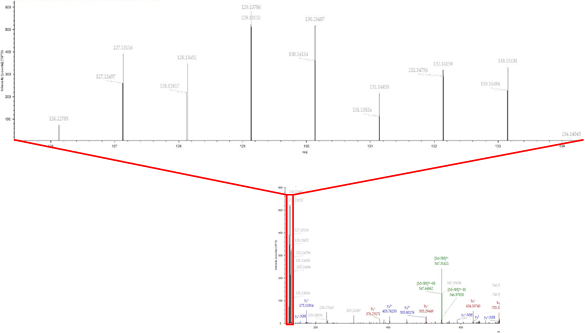
3. Label‑Free Quantitative Proteomics – (LFQ)
Scalable Quantitation for Large and Complex Studies
Our label‑free quantitative proteomics workflows enable measurement of protein abundance without chemical or isotopic labeling. This approach is well suited for large‑scale studies and complex experimental designs.
Applications
- Comparative proteomics across multiple conditions
- Biomarker discovery
- Drug development and biological pathway analysis
Label‑free workflows are supported by high‑resolution LC‑MS/MS and advanced data analysis pipelines to deliver reproducible and publication‑ready results.
4. Post‑Translational Modification (PTM) Analysis – (S60)
Comprehensive PTM Identification and Site Localization
Our PTM mass spectrometry services enable accurate identification of protein modifications using high‑resolution LC‑MS/MS workflows with confident site localization.
PTMs Analyzed
- Phosphorylation
- Acetylation and methylation
- Ubiquitination and SUMOylation
- Glycosylation
- Oxidation and other rare PTMs
"*" indicates required fields